Numpyro+jax based solution#
M. Fouesneau#
This Notebook shows how to implement a straight line fitting using MCMC and an outlier mixture model.
%%file requirements.txt
numpy
scipy
matplotlib
corner
daft
numpyro
funsor
Overwriting requirements.txt
#@title Install packages { display-mode: "form" }
# !pip install -r requirements.txt --quiet
#@title setup notebook { display-mode: "form" }
# Loading configuration
# Don't forget that mac has this annoying configuration that leads
# to limited number of figures/files
# ulimit -n 4096 <---- osx limits to 256
# Notebook matplotlib mode
%pylab inline
# set for retina or hi-resolution displays
%config InlineBackend.figure_format='retina'
light_minimal = {
'font.family': 'serif',
'font.size': 14,
"axes.titlesize": "x-large",
"axes.labelsize": "large",
'axes.edgecolor': '#666666',
"xtick.direction": "out",
"ytick.direction": "out",
"xtick.major.size": "8",
"xtick.minor.size": "4",
"ytick.major.size": "8",
"ytick.minor.size": "4",
'xtick.labelsize': 'small',
'ytick.labelsize': 'small',
'ytick.color': '#666666',
'xtick.color': '#666666',
'xtick.top': False,
'ytick.right': False,
'axes.spines.top': False,
'axes.spines.right': False,
'image.aspect': 'auto'
}
import pylab as plt
import numpy as np
plt.style.use(light_minimal)
from corner import corner
from IPython.display import display, Markdown
from itertools import cycle
from matplotlib.colors import is_color_like
def steppify(x, y):
""" Steppify a curve (x,y). Useful for manually filling histograms """
dx = 0.5 * (x[1:] + x[:-1])
xx = np.zeros( 2 * len(dx), dtype=float)
yy = np.zeros( 2 * len(y), dtype=float)
xx[0::2], xx[1::2] = dx, dx
yy[0::2], yy[1::2] = y, y
xx = np.concatenate(([x[0] - (dx[0] - x[0])], xx, [x[-1] + (x[-1] - dx[-1])]))
return xx, yy
def fast_violin(dataset, width=0.8, bins=None, color=None, which='both',
**kwargs):
""" Make a violin plot with histograms instead of kde """
if bins is None:
a, b = np.nanmin(dataset), np.nanmax(dataset)
d = np.nanstd(dataset, axis=0)
d = np.nanmin(d[d>0])
d /= np.sqrt(dataset.shape[1])
bins = np.arange(a - 1.5 * d, b + 1.5 * d, d)
default_colors = 'C0', 'C1', 'C2', 'C3', 'C4', 'C5', 'C6', 'C7', 'C8', 'C9'
if is_color_like(color):
colors = cycle([color])
elif (color is None):
colors = cycle(default_colors)
elif len(color) != dataset.shape[1]:
colors = cycle(default_colors)
else:
colors = color
for k, color_ in zip(range(dataset.shape[1]), colors):
n, bx = np.histogram(dataset[:, k], bins=bins, **kwargs)
x_ = 0.5 * (bx[1:] + bx[:-1])
y_ = n.astype(float) / n.max()
y0_ = (k + 1) + np.zeros_like(x_)
y1_ = (k + 1) + np.zeros_like(x_)
if which in ('left', 'both'):
y0_ += - 0.5 * width * y_
if which in ('right', 'both'):
y1_ += + 0.5 * width * y_
r = plt.plot(y0_, x_, color=color_)
plt.plot(y1_, x_, color=r[0].get_color())
plt.fill_betweenx(x_, y0_, y1_,
color=r[0].get_color(),
alpha=0.5)
percs_where = np.array([0.16, 0.50, 0.84])
percs_val = np.interp(percs_where, np.cumsum(y_) / np.sum(y_), x_)
plt.plot([k + 1, k + 1],
[percs_val[0], percs_val[2]], '-', color='k')
plt.plot([k + 1],
[percs_val[1]], 'o', color='k')
%pylab is deprecated, use %matplotlib inline and import the required libraries.
Populating the interactive namespace from numpy and matplotlib
Preliminary discussion around Bayesian statistics#
As astronomers, we are interested in characterizing uncertainties as much as the point-estimate in our analyses. This means we want to infer the distribution of possible/plausible parameter values.
In the Bayesian context, the Bayes’ theorem, also called chain rule of conditional probabilities, expresses how a degree of belief (prior), expressed as a probability, should rationally change to account for the availability of related evidence.
where \(A\) and \(B\) are ensembles of events. (\(P(B) \neq 0\)).
\(P(A\mid B)\) the probability of event \(A\) occurring given that an event \(B\) already occured. It is also called the posterior probability of \(A\) given \(B\).
\(P(B\mid A)\) is the probability of event \(B\) occurring given that \(A\) already occured. It is also called the likelihood of A given a fixed B.
\(P(A)\) and \(P(B)\) are the probabilities of observing \(A\) and \(B\) independently of the other one. They are known as the marginal probabilities or prior probabilities.
We note that in this equation we can swap \(A\) and \(B\). The Bayesian framework only brings interpretations. One can see it as a knowledge refinement: given what I know about \(A\) (e.g, gravity, stellar evolution, chemistry) what can we learn with new events \(B\).
A note about the proof: The equation derives from a simple observation (geometry/ensemble theory).
The figure below gives the visual support
#@title Baye's theorem figure { display-mode: "form" }
from matplotlib.patches import Ellipse, Rectangle
ax = plt.subplot(111)
patches = [
Ellipse((-1, 0), 2, 3, angle=30, facecolor='0.5', edgecolor='k', alpha=0.1),
Ellipse((-1, 0), 2, 3, angle=30, facecolor='None', edgecolor='k'),
Ellipse((1, 0), 3, 2, angle=15, facecolor='0.5', edgecolor='k', alpha=0.1),
Ellipse((1, 0), 3, 2, angle=15, facecolor='None', edgecolor='k'),
Rectangle((-3, -3), 6, 6, facecolor='None', edgecolor='k')
]
for patch in patches:
ax.add_patch(patch)
plt.text(-2.8, 2.8, r'$\Omega$', va='top', ha='left', color='k')
plt.text(-1, 0, 'A', va='top', ha='left', color='k')
plt.text(1, 0, 'B', va='top', ha='left', color='k')
plt.annotate(r'A$\cap$B', (-0.2, 0.),
xycoords='data',
arrowprops=dict(facecolor='black', arrowstyle="->"),
horizontalalignment='center', verticalalignment='bottom',
xytext=(0.5, 1.5))
plt.xlim(-3, 3)
plt.ylim(-3, 3)
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
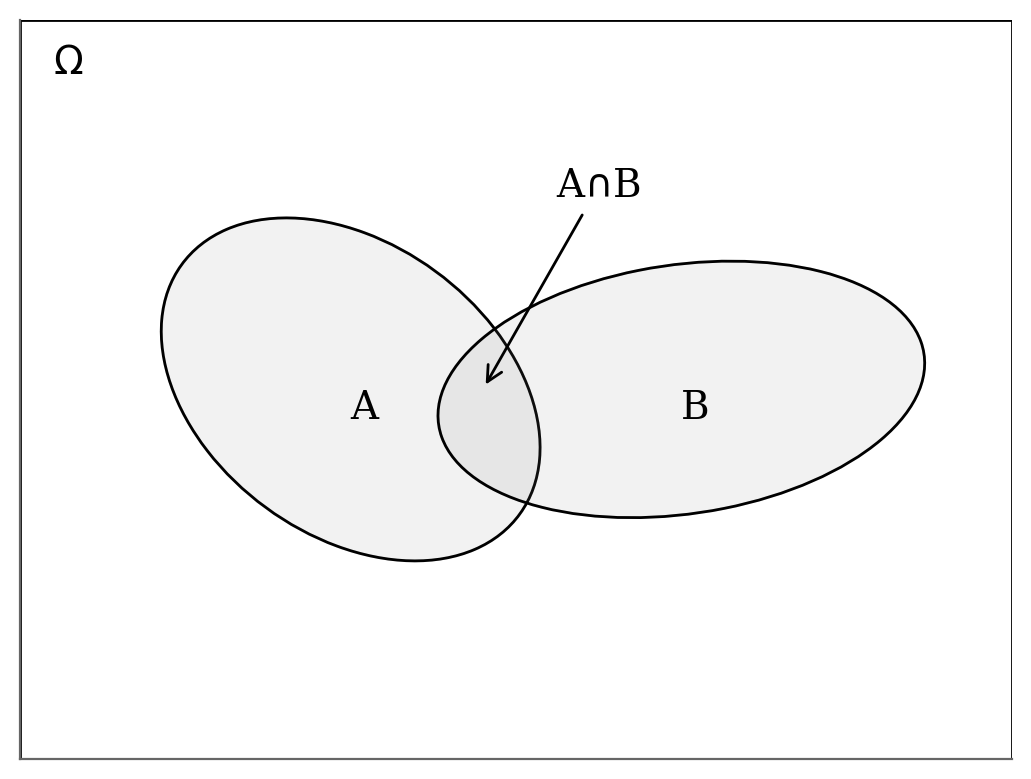
For any event \(x\) from \(\Omega\) $\( \begin{eqnarray} P(x\in A \cap B) & = P(x\in A \mid x \in B) \cdot P(x\in B)\\ & = P(x\in B \mid x \in A) \cdot P(x\in A) \end{eqnarray} \)\( which we can write in a more compact way as \)\( \begin{eqnarray} P(A, B \mid \Omega) & = P(A \mid B, \Omega) \cdot P(B \mid \Omega)\\ & = P(B \mid A, \Omega) \cdot P(A \mid \Omega)\\ \end{eqnarray} \)\( Note that, we explicitly noted \)\Omega$ but it is commonly omitted. The final equation of the Bayes’ theorem derives from the rearanging the terms of the right hand-side equalities.
The equation itself is most commonly attributed to reverand Thomas Bayes in 1763, but also to Pierre-Simon de Laplace. However, the current interpretation and usage significantly expanded from its initial formulation.
Straight line problem#
In astronomy, we commonly have to fit a model to data. A very typical example is the fit of a straight line to a set of points in a two-dimensional plane. Astronomers are very good at finding a representation or a transformation of their data in which they can identify linear correlations (e.g., plotting in log-log space, action-angles for orbits, etc). Consequently, a polynomial remains a “line” in a more complicated representation space.
In this exercise, we consider a dataset with uncertainties along one axis only for simplicity, i.e. uncertainties along x negligible. These conditions are rarely met in practice but the principles remain similar. However, we also explore the outlier problem: bad data are very common in astronomy.
We emphasize the importance of having a “generative model” for the data, even an approximate one. Once there is a generative model, we have a direct computation of the likelihood of the parameters or the posterior probability distribution.
Problem definition#
We will fit a linear model to a dataset \(\{(x_i, y_i, \sigma_i)\}\)
Let’s start by looking at the data.
Generate dataset: Straight line with outliers#
#@title Generate data { display-mode: "form" }
x = np.array([ 0, 3, 9, 14, 15, 19, 20, 21, 30, 35,
40, 41, 42, 43, 54, 56, 67, 69, 72, 88])
y = np.array([33, 68, 34, 34, 37, 71, 37, 44, 48, 49,
53, 49, 50, 48, 56, 60, 61, 63, 44, 71])
sy = np.array([ 3.6, 3.9, 2.6, 3.4, 3.8, 3.8, 2.2, 2.1, 2.3, 3.8,
2.2, 2.8, 3.9, 3.1, 3.4, 2.6, 3.4, 3.7, 2.0, 3.5])
np.savetxt('line_outlier.dat', np.array([x, y, sy]).T)
x, y, sy = np.loadtxt('line_outlier.dat', unpack=True)
plt.errorbar(x, y, yerr=sy, marker='o', ls='None', ms=5, color='k');
plt.xlabel('X');
plt.ylabel('Y');
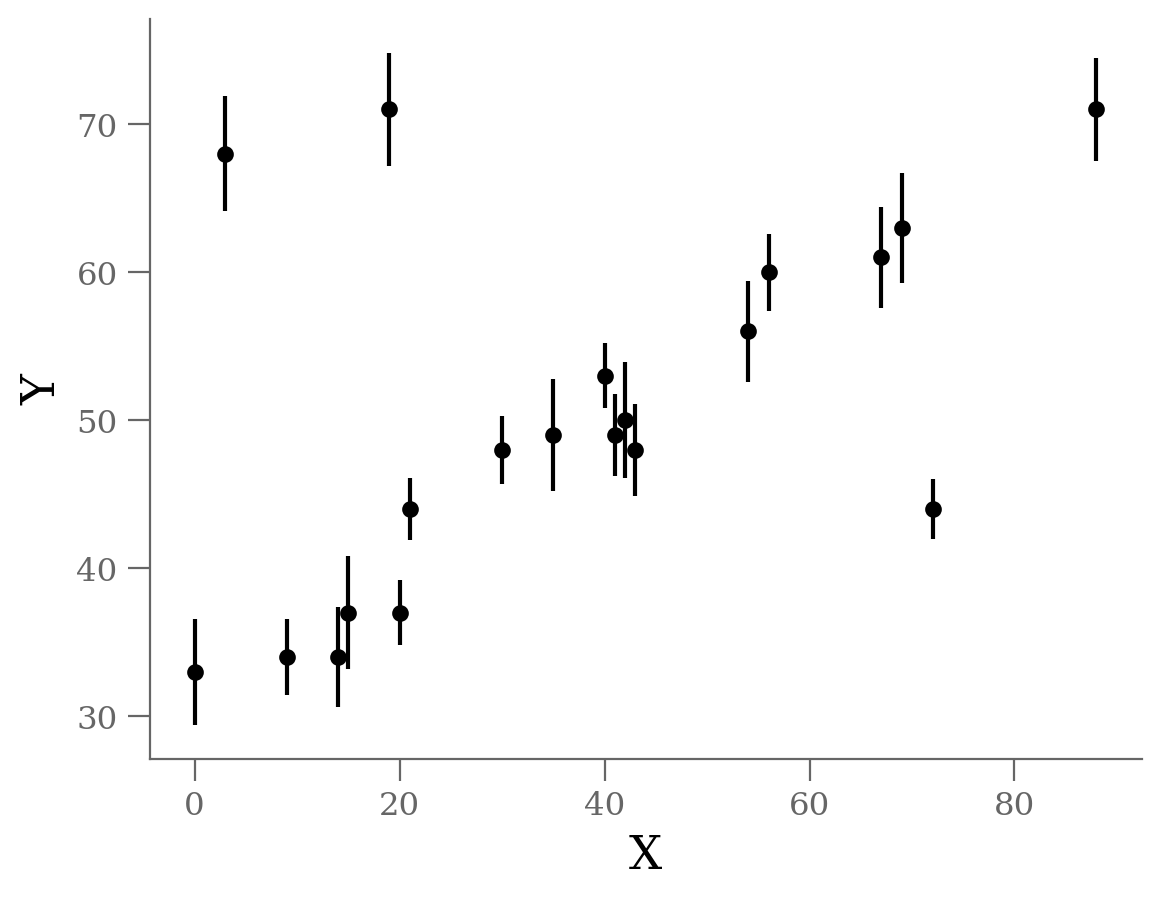
We see three outliers, but there is an obvious linear relation between the points overall.
Blind fit: no outlier model#
Let’s see what happens if we fit the data without any outlier consideration.
Equations#
We would like to fit a linear model to this above dataset:
We commonly start with the Bayes’s rule:
Hence we need to define our prior and likelihood
Given this model and the gaussian uncertainties on our data, we can compute a Gaussian likelihood for each point: $\( P(x_i,y_i,~|~\alpha, \beta, \sigma_i) = \frac{1}{\sqrt{2\pi\sigma_i^2}} \exp\left[-\frac{1}{2\sigma_i^2}\left(y_i - {y}(x_i~|~\alpha, \beta)\right)^2\right] \)\( Note that \)\sigma_i\( is on the right hand side of the \)\mid$. It is because we assume given the Gaussian uncertainties when we write the likelihood that way.
The total likelihood is the product of all the individual likelihoods (as we assume independent measurements).
For numerical stability reasons, it is preferable to take the log-likelihood of the data. We have: $\( \log P(\{x_i, y_i\}~|~\alpha, \beta, \{\sigma_i\}) = \mathrm{const} - \sum_i \frac{1}{2\sigma_i^2}\left(y_i - y(x_i~|~\alpha, \beta)\right)^2 \)$
The posterior is the likelihood times the prior distributions. Let’s assume from the inspection of the data that \(\alpha > 0\), we can write $\(P(\alpha) = Uniform[0, 100], \)\( and \)\(P(\beta) = Uniform[0, 100].\)$ (The upper value is abitrary.)
Coding#
JAX is Autograd and XLA, brought together for high-performance machine learning research supported by Google.
Autograd: automatically differentiate native Python and NumPy functions. It can differentiate through loops, branches, recursion, and closures, and it can take derivatives of derivatives of derivatives. It supports reverse-mode differentiation (a.k.a. backpropagation) via grad as well as forward-mode differentiation, and the two can be composed arbitrarily to any order.
XLA: compile and run your NumPy programs on GPUs and TPUs. Compilation happens under the hood by default, with library calls getting just-in-time compiled and executed.
import jax.numpy as jnp
from jax import random, vmap
from jax.scipy.special import logsumexp
import numpyro
from numpyro.diagnostics import hpdi
import numpyro.distributions as dist
from numpyro import handlers
from numpyro.infer import MCMC, NUTS
/opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
def jax_model_ols(x=None, y=None, sigma_y=None):
beta = numpyro.sample("beta", dist.Uniform(0.0, 100))
alpha = numpyro.sample("alpha", dist.Uniform(0.0, 100))
ypred = beta + alpha * x
numpyro.sample("y", dist.Normal(ypred, sigma_y), obs=y)
numpyro.render_model(jax_model_ols, model_args=(x, y, sy,),
render_distributions=True)
---------------------------------------------------------------------------
FileNotFoundError Traceback (most recent call last)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:76, in run_check(cmd, input_lines, encoding, quiet, **kwargs)
75 kwargs['stdout'] = kwargs['stderr'] = subprocess.PIPE
---> 76 proc = _run_input_lines(cmd, input_lines, kwargs=kwargs)
77 else:
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:96, in _run_input_lines(cmd, input_lines, kwargs)
95 def _run_input_lines(cmd, input_lines, *, kwargs):
---> 96 popen = subprocess.Popen(cmd, stdin=subprocess.PIPE, **kwargs)
98 stdin_write = popen.stdin.write
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/subprocess.py:858, in Popen.__init__(self, args, bufsize, executable, stdin, stdout, stderr, preexec_fn, close_fds, shell, cwd, env, universal_newlines, startupinfo, creationflags, restore_signals, start_new_session, pass_fds, encoding, errors, text)
855 self.stderr = io.TextIOWrapper(self.stderr,
856 encoding=encoding, errors=errors)
--> 858 self._execute_child(args, executable, preexec_fn, close_fds,
859 pass_fds, cwd, env,
860 startupinfo, creationflags, shell,
861 p2cread, p2cwrite,
862 c2pread, c2pwrite,
863 errread, errwrite,
864 restore_signals, start_new_session)
865 except:
866 # Cleanup if the child failed starting.
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/subprocess.py:1720, in Popen._execute_child(self, args, executable, preexec_fn, close_fds, pass_fds, cwd, env, startupinfo, creationflags, shell, p2cread, p2cwrite, c2pread, c2pwrite, errread, errwrite, restore_signals, start_new_session)
1719 err_msg = os.strerror(errno_num)
-> 1720 raise child_exception_type(errno_num, err_msg, err_filename)
1721 raise child_exception_type(err_msg)
FileNotFoundError: [Errno 2] No such file or directory: PosixPath('dot')
The above exception was the direct cause of the following exception:
ExecutableNotFound Traceback (most recent call last)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/IPython/core/formatters.py:974, in MimeBundleFormatter.__call__(self, obj, include, exclude)
971 method = get_real_method(obj, self.print_method)
973 if method is not None:
--> 974 return method(include=include, exclude=exclude)
975 return None
976 else:
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:98, in JupyterIntegration._repr_mimebundle_(self, include, exclude, **_)
96 include = set(include) if include is not None else {self._jupyter_mimetype}
97 include -= set(exclude or [])
---> 98 return {mimetype: getattr(self, method_name)()
99 for mimetype, method_name in MIME_TYPES.items()
100 if mimetype in include}
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:98, in <dictcomp>(.0)
96 include = set(include) if include is not None else {self._jupyter_mimetype}
97 include -= set(exclude or [])
---> 98 return {mimetype: getattr(self, method_name)()
99 for mimetype, method_name in MIME_TYPES.items()
100 if mimetype in include}
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:112, in JupyterIntegration._repr_image_svg_xml(self)
110 def _repr_image_svg_xml(self) -> str:
111 """Return the rendered graph as SVG string."""
--> 112 return self.pipe(format='svg', encoding=SVG_ENCODING)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:104, in Pipe.pipe(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
55 def pipe(self,
56 format: typing.Optional[str] = None,
57 renderer: typing.Optional[str] = None,
(...)
61 engine: typing.Optional[str] = None,
62 encoding: typing.Optional[str] = None) -> typing.Union[bytes, str]:
63 """Return the source piped through the Graphviz layout command.
64
65 Args:
(...)
102 '<?xml version='
103 """
--> 104 return self._pipe_legacy(format,
105 renderer=renderer,
106 formatter=formatter,
107 neato_no_op=neato_no_op,
108 quiet=quiet,
109 engine=engine,
110 encoding=encoding)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/_tools.py:171, in deprecate_positional_args.<locals>.decorator.<locals>.wrapper(*args, **kwargs)
162 wanted = ', '.join(f'{name}={value!r}'
163 for name, value in deprecated.items())
164 warnings.warn(f'The signature of {func.__name__} will be reduced'
165 f' to {supported_number} positional args'
166 f' {list(supported)}: pass {wanted}'
167 ' as keyword arg(s)',
168 stacklevel=stacklevel,
169 category=category)
--> 171 return func(*args, **kwargs)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:121, in Pipe._pipe_legacy(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
112 @_tools.deprecate_positional_args(supported_number=2)
113 def _pipe_legacy(self,
114 format: typing.Optional[str] = None,
(...)
119 engine: typing.Optional[str] = None,
120 encoding: typing.Optional[str] = None) -> typing.Union[bytes, str]:
--> 121 return self._pipe_future(format,
122 renderer=renderer,
123 formatter=formatter,
124 neato_no_op=neato_no_op,
125 quiet=quiet,
126 engine=engine,
127 encoding=encoding)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:149, in Pipe._pipe_future(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
146 if encoding is not None:
147 if codecs.lookup(encoding) is codecs.lookup(self.encoding):
148 # common case: both stdin and stdout need the same encoding
--> 149 return self._pipe_lines_string(*args, encoding=encoding, **kwargs)
150 try:
151 raw = self._pipe_lines(*args, input_encoding=self.encoding, **kwargs)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/piping.py:212, in pipe_lines_string(engine, format, input_lines, encoding, renderer, formatter, neato_no_op, quiet)
206 cmd = dot_command.command(engine, format,
207 renderer=renderer,
208 formatter=formatter,
209 neato_no_op=neato_no_op)
210 kwargs = {'input_lines': input_lines, 'encoding': encoding}
--> 212 proc = execute.run_check(cmd, capture_output=True, quiet=quiet, **kwargs)
213 return proc.stdout
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:81, in run_check(cmd, input_lines, encoding, quiet, **kwargs)
79 except OSError as e:
80 if e.errno == errno.ENOENT:
---> 81 raise ExecutableNotFound(cmd) from e
82 raise
84 if not quiet and proc.stderr:
ExecutableNotFound: failed to execute PosixPath('dot'), make sure the Graphviz executables are on your systems' PATH
<graphviz.graphs.Digraph at 0x7fa7a34a4d00>
def jax_model_ols(x, y, sigma_y):
beta = numpyro.sample("beta", dist.Normal(0.0, 100))
alpha = numpyro.sample("alpha", dist.Normal(0.0, 100))
with numpyro.plate("i=1..n", len(y)):
ypred = beta + alpha * x
numpyro.sample("y", dist.Normal(ypred, sigma_y), obs=y)
numpyro.render_model(jax_model_ols, model_args=(x, y, sy,),
render_distributions=True)
---------------------------------------------------------------------------
FileNotFoundError Traceback (most recent call last)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:76, in run_check(cmd, input_lines, encoding, quiet, **kwargs)
75 kwargs['stdout'] = kwargs['stderr'] = subprocess.PIPE
---> 76 proc = _run_input_lines(cmd, input_lines, kwargs=kwargs)
77 else:
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:96, in _run_input_lines(cmd, input_lines, kwargs)
95 def _run_input_lines(cmd, input_lines, *, kwargs):
---> 96 popen = subprocess.Popen(cmd, stdin=subprocess.PIPE, **kwargs)
98 stdin_write = popen.stdin.write
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/subprocess.py:858, in Popen.__init__(self, args, bufsize, executable, stdin, stdout, stderr, preexec_fn, close_fds, shell, cwd, env, universal_newlines, startupinfo, creationflags, restore_signals, start_new_session, pass_fds, encoding, errors, text)
855 self.stderr = io.TextIOWrapper(self.stderr,
856 encoding=encoding, errors=errors)
--> 858 self._execute_child(args, executable, preexec_fn, close_fds,
859 pass_fds, cwd, env,
860 startupinfo, creationflags, shell,
861 p2cread, p2cwrite,
862 c2pread, c2pwrite,
863 errread, errwrite,
864 restore_signals, start_new_session)
865 except:
866 # Cleanup if the child failed starting.
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/subprocess.py:1720, in Popen._execute_child(self, args, executable, preexec_fn, close_fds, pass_fds, cwd, env, startupinfo, creationflags, shell, p2cread, p2cwrite, c2pread, c2pwrite, errread, errwrite, restore_signals, start_new_session)
1719 err_msg = os.strerror(errno_num)
-> 1720 raise child_exception_type(errno_num, err_msg, err_filename)
1721 raise child_exception_type(err_msg)
FileNotFoundError: [Errno 2] No such file or directory: PosixPath('dot')
The above exception was the direct cause of the following exception:
ExecutableNotFound Traceback (most recent call last)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/IPython/core/formatters.py:974, in MimeBundleFormatter.__call__(self, obj, include, exclude)
971 method = get_real_method(obj, self.print_method)
973 if method is not None:
--> 974 return method(include=include, exclude=exclude)
975 return None
976 else:
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:98, in JupyterIntegration._repr_mimebundle_(self, include, exclude, **_)
96 include = set(include) if include is not None else {self._jupyter_mimetype}
97 include -= set(exclude or [])
---> 98 return {mimetype: getattr(self, method_name)()
99 for mimetype, method_name in MIME_TYPES.items()
100 if mimetype in include}
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:98, in <dictcomp>(.0)
96 include = set(include) if include is not None else {self._jupyter_mimetype}
97 include -= set(exclude or [])
---> 98 return {mimetype: getattr(self, method_name)()
99 for mimetype, method_name in MIME_TYPES.items()
100 if mimetype in include}
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/jupyter_integration.py:112, in JupyterIntegration._repr_image_svg_xml(self)
110 def _repr_image_svg_xml(self) -> str:
111 """Return the rendered graph as SVG string."""
--> 112 return self.pipe(format='svg', encoding=SVG_ENCODING)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:104, in Pipe.pipe(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
55 def pipe(self,
56 format: typing.Optional[str] = None,
57 renderer: typing.Optional[str] = None,
(...)
61 engine: typing.Optional[str] = None,
62 encoding: typing.Optional[str] = None) -> typing.Union[bytes, str]:
63 """Return the source piped through the Graphviz layout command.
64
65 Args:
(...)
102 '<?xml version='
103 """
--> 104 return self._pipe_legacy(format,
105 renderer=renderer,
106 formatter=formatter,
107 neato_no_op=neato_no_op,
108 quiet=quiet,
109 engine=engine,
110 encoding=encoding)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/_tools.py:171, in deprecate_positional_args.<locals>.decorator.<locals>.wrapper(*args, **kwargs)
162 wanted = ', '.join(f'{name}={value!r}'
163 for name, value in deprecated.items())
164 warnings.warn(f'The signature of {func.__name__} will be reduced'
165 f' to {supported_number} positional args'
166 f' {list(supported)}: pass {wanted}'
167 ' as keyword arg(s)',
168 stacklevel=stacklevel,
169 category=category)
--> 171 return func(*args, **kwargs)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:121, in Pipe._pipe_legacy(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
112 @_tools.deprecate_positional_args(supported_number=2)
113 def _pipe_legacy(self,
114 format: typing.Optional[str] = None,
(...)
119 engine: typing.Optional[str] = None,
120 encoding: typing.Optional[str] = None) -> typing.Union[bytes, str]:
--> 121 return self._pipe_future(format,
122 renderer=renderer,
123 formatter=formatter,
124 neato_no_op=neato_no_op,
125 quiet=quiet,
126 engine=engine,
127 encoding=encoding)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/piping.py:149, in Pipe._pipe_future(self, format, renderer, formatter, neato_no_op, quiet, engine, encoding)
146 if encoding is not None:
147 if codecs.lookup(encoding) is codecs.lookup(self.encoding):
148 # common case: both stdin and stdout need the same encoding
--> 149 return self._pipe_lines_string(*args, encoding=encoding, **kwargs)
150 try:
151 raw = self._pipe_lines(*args, input_encoding=self.encoding, **kwargs)
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/piping.py:212, in pipe_lines_string(engine, format, input_lines, encoding, renderer, formatter, neato_no_op, quiet)
206 cmd = dot_command.command(engine, format,
207 renderer=renderer,
208 formatter=formatter,
209 neato_no_op=neato_no_op)
210 kwargs = {'input_lines': input_lines, 'encoding': encoding}
--> 212 proc = execute.run_check(cmd, capture_output=True, quiet=quiet, **kwargs)
213 return proc.stdout
File /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/graphviz/backend/execute.py:81, in run_check(cmd, input_lines, encoding, quiet, **kwargs)
79 except OSError as e:
80 if e.errno == errno.ENOENT:
---> 81 raise ExecutableNotFound(cmd) from e
82 raise
84 if not quiet and proc.stderr:
ExecutableNotFound: failed to execute PosixPath('dot'), make sure the Graphviz executables are on your systems' PATH
<graphviz.graphs.Digraph at 0x7fa7a3473400>
# Start from this source of randomness. We will split keys for subsequent operations.
rng_key = random.PRNGKey(0)
rng_key, rng_key_ = random.split(rng_key)
# Run NUTS.
kernel = NUTS(jax_model_ols)
num_samples = 5000
mcmc = MCMC(kernel, num_warmup=3000, num_samples=num_samples)
mcmc.run(rng_key_, x=x, y=y, sigma_y=sy)
mcmc.print_summary()
traces_ols = mcmc.get_samples()
No GPU/TPU found, falling back to CPU. (Set TF_CPP_MIN_LOG_LEVEL=0 and rerun for more info.)
mean std median 5.0% 95.0% n_eff r_hat
alpha 0.24 0.03 0.24 0.19 0.28 1398.30 1.00
beta 39.72 1.24 39.72 37.68 41.78 1390.04 1.00
Number of divergences: 0
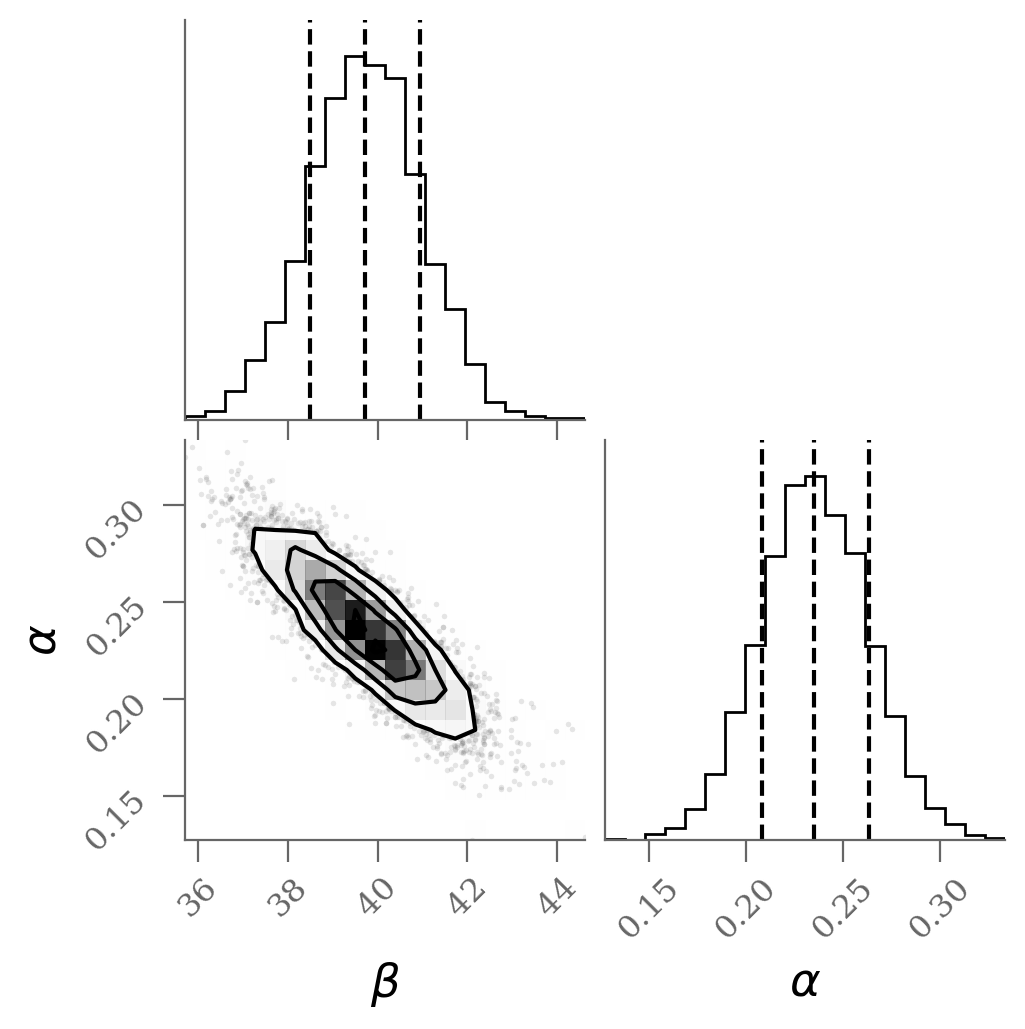
varnames = 'beta', 'alpha'
sample = np.array([traces_ols[k].ravel() for k in varnames]).T
corner(sample, labels=(r'$\beta$', r'$\alpha$'), quantiles=(0.16, 0.50, 0.84));
x, y, sy = np.loadtxt('line_outlier.dat', unpack=True)
plt.errorbar(x, y, yerr=sy, marker='o', ls='None', ms=5, color='k');
plt.xlabel('X');
plt.ylabel('Y');
ypred_blind = sample[:, 0] + sample[:, 1] * x[:, None]
plt.plot(x, ypred_blind[:, :100], color='C0', rasterized=True, alpha=0.3, lw=0.3)
percs_blind = np.percentile(sample, [16, 50, 84], axis=0)
display(Markdown(r"""Without outlier modeling:
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
""".format(*percs_blind.T.ravel())
))
Without outlier modeling:
$\alpha$ = 39.7 [38.5, 40.9]
$\beta$ = 0.235 [0.208, 0.263]
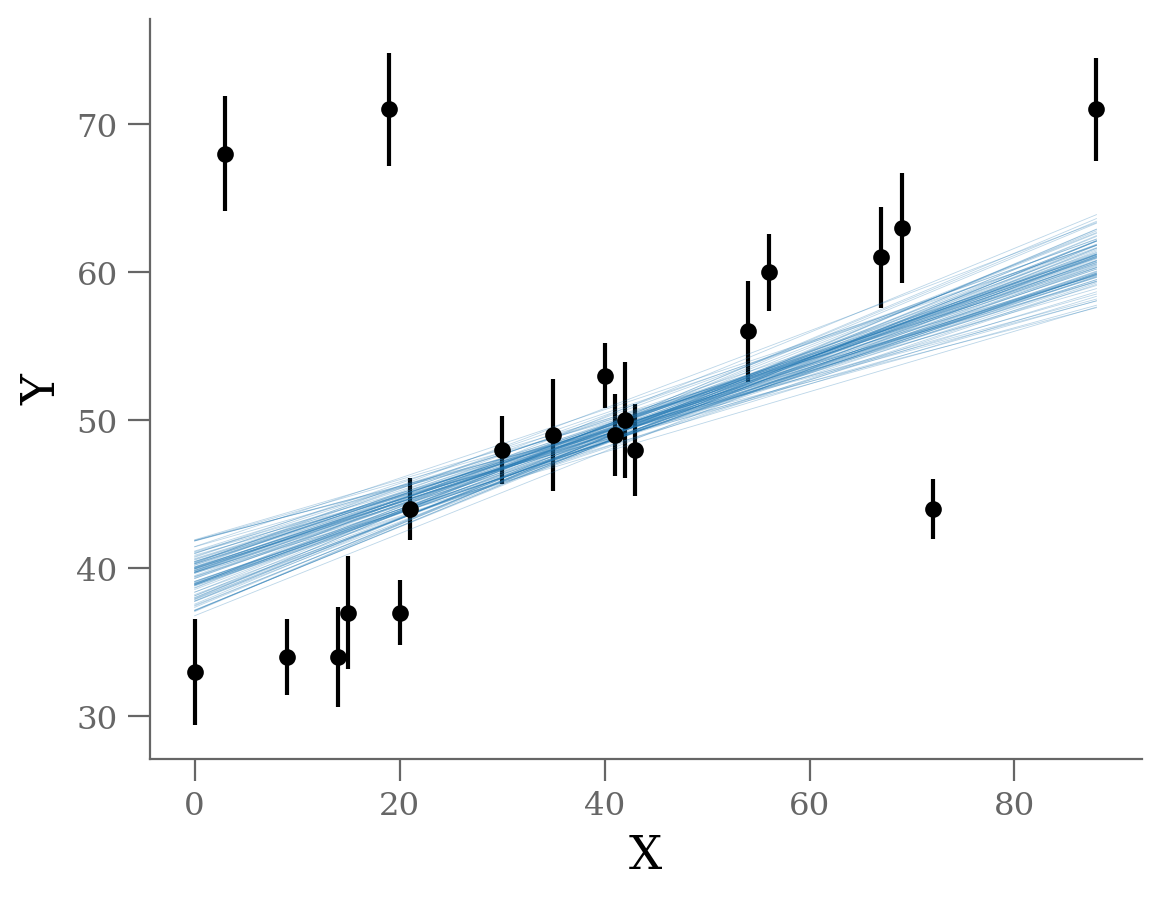
It’s clear from this plot that the outliers exerts a disproportionate influence on the fit.
This is due to the nature of our likelihood function. One outlier that is, say \(10-\sigma\) (standard deviations) away from the fit will out-weight the contribution of points which are only \(2-\sigma\) away.
In conclusion, least-square likelihoods are overly sensitive to outliers, and this is causing issues with our fit.
One way to address this is to simply model the outliers.
Mixture Model#
The Bayesian approach to accounting for outliers generally involves mixture models so that the initial model is combined with a complement model accounting for the outliers:
where \(q\) is the mixing coefficient (between 0 and 1), \(P_{in}\) and \(P_{out}\) the probabilities of being an inlier and outlier, respectively.
So let’s propose a more complicated model that is a mixture between a signal and a background
Graphical model#
A short parenthesis on probabilistic programming. One way to represent the above model is through probabilistic graphical models (pgm).
Using daft a python package, I draw a (inaccurate) pgm of the straight line model with outliers.
import daft
# Instantiate the PGM.
pgm = daft.PGM([4, 4], origin=[-0.5, -0.0], grid_unit=4,
node_unit=2, label_params=dict(fontsize='x-large'))
# Hierarchical parameters.
pgm.add_node(daft.Node("sout", r"$\sigma_{out}$", 0.9, 3.5, fixed=False))
pgm.add_node(daft.Node("g", r"$g$", 0.2, 3.5, fixed=False))
pgm.add_node(daft.Node("beta", r"$\beta$", 2.4, 3.5))
pgm.add_node(daft.Node("alpha", r"$\alpha$", 1.6, 3.5))
# Latent variable.
pgm.add_node(daft.Node("x", r"$x_i$", 1, 1))
pgm.add_node(daft.Node("gn", r"$g_i$", 0.2, 2, observed=False))
pgm.add_node(daft.Node("yin", r"$\hat{y}_{in}$", 2.5, 2, observed=False))
pgm.add_node(daft.Node("yout", r"$\hat{y}_{out}$", 1.5, 2, observed=False))
# Data.
pgm.add_node(daft.Node("y", r"$y_i$", 2, 1, observed=True))
# And a plate.
pgm.add_plate(daft.Plate([-0.2, 0.5, 3.2, 2.2],
label=r"$i = 1, \cdots, N$", shift=-0.1,
bbox={"color": "none"}))
# add relations
pgm.add_edge("x", "y")
pgm.add_edge("alpha", "yin")
pgm.add_edge("beta", "yin")
pgm.add_edge("alpha", "yout")
pgm.add_edge("beta", "yout")
pgm.add_edge("sout", "yout")
pgm.add_edge("g", "gn")
pgm.add_edge("gn", "y")
pgm.add_edge("yin", "y")
pgm.add_edge("yout", "y")
pgm.render();
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
Cell In[13], line 1
----> 1 import daft
3 # Instantiate the PGM.
4 pgm = daft.PGM([4, 4], origin=[-0.5, -0.0], grid_unit=4,
5 node_unit=2, label_params=dict(fontsize='x-large'))
ModuleNotFoundError: No module named 'daft'
There are multiple ways to define this mixture modeling.
Brutal version: 1 parameter per datapoint#
\(P_{in, i}\) corresponds to the previous likelihood.
\(P_{out, i}\) is arbitrary as we do not really have information about the causes for the outliers. We assume a similar Gaussian form centered on the affine relation but with a significantly larger dispersion. Doing so is intuitively considering the distance to the line to decide whether we have an outlier or not (this is intuitively a Bayesian version of sigma-clipping)
It is important to note that the total likelihood is a product of sums:
$\( P = \prod_i P_i = \prod_i \left(q_i \cdot P_{in, i} + (1 - q_i) \cdot P_{out, i}\right).\)\(
Hence, we need to be careful to properly transform the \)\ln P\( into \)P$ during the calculations (see: np.logsumexp)
Hence the likelihood becomes explicitly $\( \begin{array}{ll} P(\{x_i\}, \{y_i\}~|~\theta,\{q_i\},\{\sigma_i\}, \sigma_{out}) = & \frac{q_i}{\sqrt{2\pi \sigma_i^2}}\exp\left[\frac{-\left(\hat{y}(x_i~|~\theta) - y_i\right)^2}{2\sigma_i^2}\right] \\ &+ \frac{1 - q_i}{\sqrt{2\pi \sigma_{out}^2}}\exp\left[\frac{-\left(\hat{y}(x_i~|~\theta) - y_i\right)^2}{2\sigma_{out}^2}\right]. \end{array} \)$
We “simply” expanded our model with \(20\) nuisance parameters: \(\{q_i\}\) is a series of weights, which range from 0 to 1 and encode for each point \(i\) the degree to which it fits the model.
\(q_i=0\) indicates an outlier, in which case a Gaussian of width \(\sigma_{out}\) is used in the computation of the likelihood. This \(\sigma_{out}\) can also be a nuisance parameter. However, for this example, we fix \(\sigma_{out}\) to an arbitrary large value (large compared with the dispersion of the data)
Note that we have a prior that all \(q_i\) must be strictly between 0 and 1.
We now make the probabilistic equations that link the various variables together:
We would like to fit a linear model to this above dataset:
One must provide some prior to properly implement the model. Let’s suppose weak informative normal priors (they correspond to a Ridge regression context)
We also have the outlier component \(\hat{y}_{out}\) $\( \hat{y}_{out} \rightsquigarrow \mathcal{N}(\hat{y}_{out}, \sigma_{out})\)\( where \)\sigma_{out}\( is unknown but positively constrained (half-Normal prior) \)\( \sigma_{out} \rightsquigarrow \mathcal{N^+}(0, 100)\)$
And each data point has a probability (0 or 1) to be an outlier or not with a probability \(g_i\) $\( g_i \rightsquigarrow \mathcal{U}(0, 1) \)$ (this is of course, sub-optimal)
Our final model is a mixture of the two components:
We code the model as:
def jax_model_brute(x=None, y=None, sigma_y=None):
## Define weakly informative Normal priors
beta = numpyro.sample("beta", dist.Uniform(0.0, 100))
alpha = numpyro.sample("alpha", dist.Uniform(0.0, 100))
## variance of outliers
sigma_y_out = numpyro.sample("sigma_y_out", dist.HalfNormal(100))
## model
ypred = beta + alpha * x
with numpyro.plate("i=1..n", len(y), dim=-1):
## define the linear model
p_outlier = numpyro.sample('p_outlier',
dist.Uniform(0, 1))
## need a float for the mixture
probs = jnp.stack([1 - p_outlier, p_outlier], axis=-1)
locs_ = jnp.stack([ypred, ypred], -1)
scales_ = jnp.stack([sigma_y, sigma_y_out * jnp.ones(len(x))], -1)
comp_ = dist.Normal(locs_, scales_)
mixture = dist.MixtureSameFamily(dist.Categorical(probs), comp_)
# likelihood
numpyro.sample("y", mixture, obs=y)
numpyro.render_model(jax_model_brute, model_args=(x, y, sy,),
render_distributions=True)
# Start from this source of randomness. We will split keys for subsequent operations.
rng_key = random.PRNGKey(0)
rng_key, rng_key_ = random.split(rng_key)
# Run NUTS.
kernel = NUTS(jax_model_brute)
num_samples = 10_000
mcmc = MCMC(kernel, num_warmup=50_000, num_samples=num_samples)
mcmc.run(rng_key_, x=x, y=y, sigma_y=sy)
mcmc.print_summary()
traces = mcmc.get_samples()
sample: 100%|██████████| 60000/60000 [00:33<00:00, 1772.17it/s, 15 steps of size 3.20e-01. acc. prob=0.92]
mean std median 5.0% 95.0% n_eff r_hat
alpha 0.46 0.04 0.46 0.39 0.53 10480.49 1.00
beta 31.74 1.89 31.67 28.53 34.68 10084.71 1.00
p_outlier[0] 0.38 0.26 0.34 0.00 0.78 19142.27 1.00
p_outlier[1] 0.67 0.24 0.71 0.31 1.00 20917.21 1.00
p_outlier[2] 0.38 0.27 0.34 0.00 0.79 16707.34 1.00
p_outlier[3] 0.42 0.28 0.38 0.00 0.83 16934.32 1.00
p_outlier[4] 0.39 0.27 0.35 0.00 0.79 18279.33 1.00
p_outlier[5] 0.67 0.23 0.70 0.32 1.00 18118.53 1.00
p_outlier[6] 0.45 0.29 0.42 0.00 0.87 16598.04 1.00
p_outlier[7] 0.40 0.27 0.36 0.00 0.80 17744.98 1.00
p_outlier[8] 0.39 0.27 0.34 0.00 0.79 20899.22 1.00
p_outlier[9] 0.38 0.26 0.34 0.00 0.78 18410.61 1.00
p_outlier[10] 0.40 0.27 0.35 0.00 0.80 18262.61 1.00
p_outlier[11] 0.38 0.26 0.33 0.00 0.77 18054.81 1.00
p_outlier[12] 0.38 0.26 0.34 0.00 0.78 17200.75 1.00
p_outlier[13] 0.40 0.27 0.36 0.00 0.81 19761.69 1.00
p_outlier[14] 0.38 0.26 0.34 0.00 0.77 17896.72 1.00
p_outlier[15] 0.39 0.26 0.35 0.00 0.79 19058.08 1.00
p_outlier[16] 0.38 0.27 0.34 0.00 0.79 19299.24 1.00
p_outlier[17] 0.38 0.26 0.34 0.00 0.76 17660.67 1.00
p_outlier[18] 0.66 0.23 0.70 0.32 1.00 22792.29 1.00
p_outlier[19] 0.39 0.27 0.35 0.00 0.80 17606.22 1.00
sigma_y_out 28.82 15.12 24.67 12.34 46.89 6879.70 1.00
Number of divergences: 0
varnames = 'beta', 'alpha', 'sigma_y_out'
sample = np.array([traces[k].ravel() for k in varnames]).T
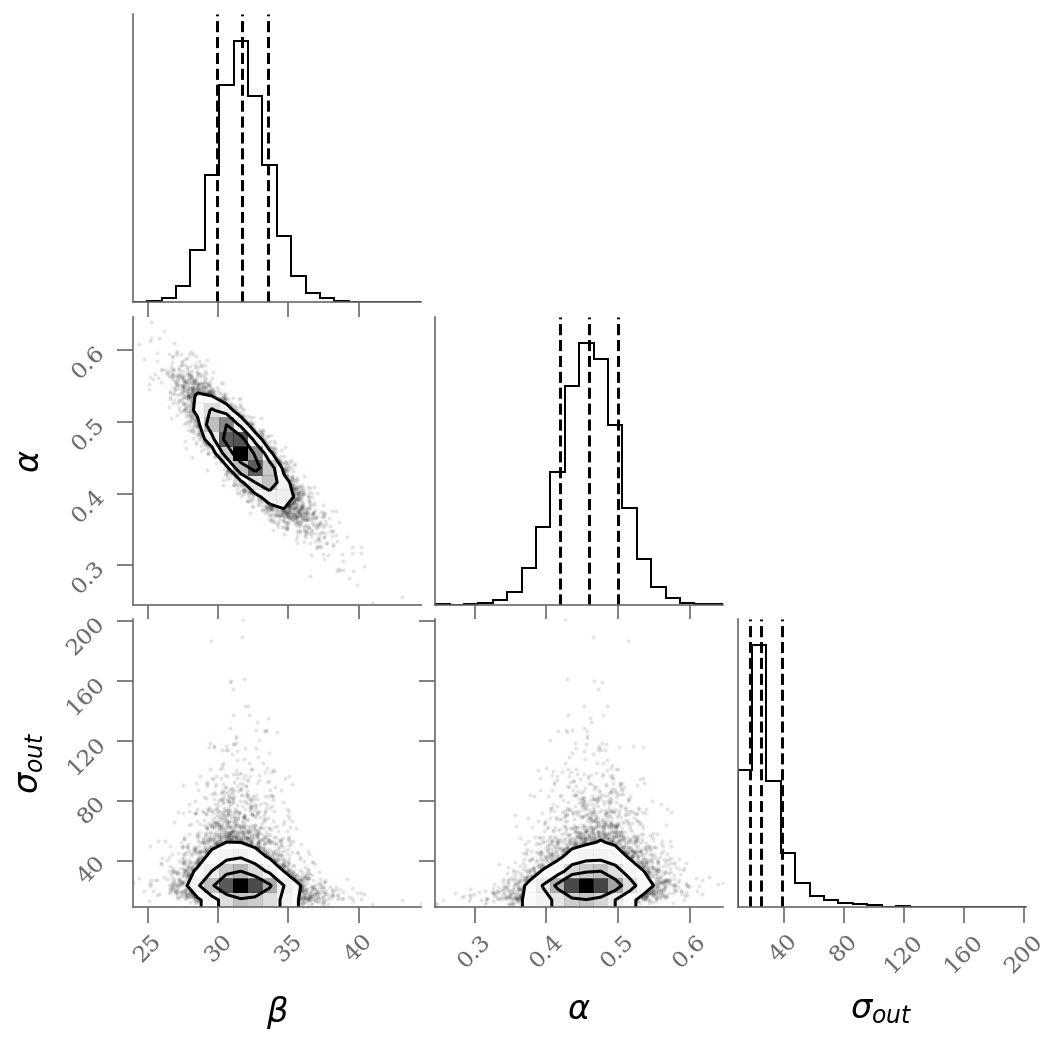
corner(sample[:, :], labels=(r'$\beta$', r'$\alpha$', r'$\sigma_{out}$'),
quantiles=(0.16, 0.50, 0.84));
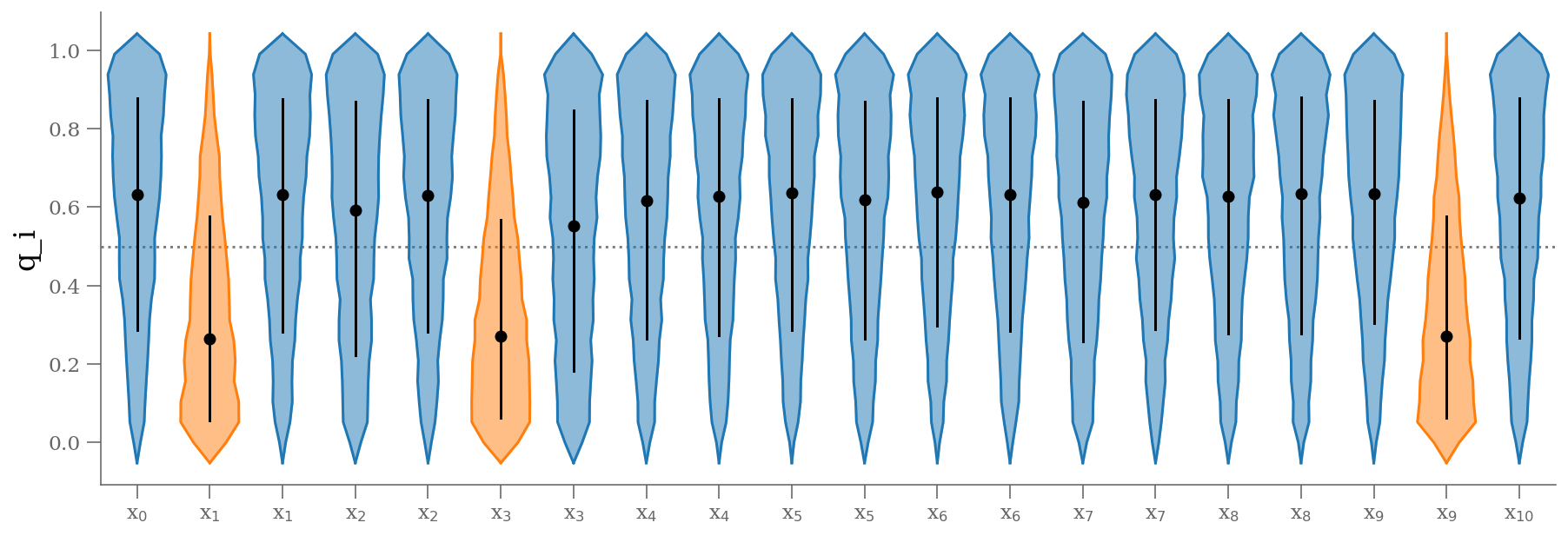
samples_q = 1 - traces['p_outlier'].T
plt.figure(figsize=(15, 5))
colors = np.where(np.median(samples_q, axis=1) < 0.5, 'C1', 'C0')
fast_violin(samples_q.T, color=colors)
ticks = (np.arange(samples_q.shape[0]) + 1)
lbls = [r'x$_{{{0:d}}}$'.format(k) for k in (0.5 * ticks).astype(int)]
plt.xticks(ticks, lbls)
plt.hlines([0.5], 0.5, len(ticks) + 0.5, color='0.5', zorder=-100, ls=':')
plt.xlim(0.5, len(ticks) + 0.5)
plt.ylabel('q_i');
q = np.median(samples_q, 1)
outliers = (q <= 0.5) # arbitrary choice
n_outliers = sum(outliers)
x, y, sy = np.loadtxt('line_outlier.dat', unpack=True)
plt.errorbar(x, y, yerr=sy, marker='o', ls='None', ms=5, color='k');
ypred_brute = sample[:, 0] + sample[:, 1] * x[:, None]
plt.plot(x, ypred_blind[:, :100], color='C0', rasterized=True, alpha=0.3, lw=0.3)
plt.plot(x, ypred_brute[:, :100], color='C1', rasterized=True, alpha=0.3, lw=0.3)
plt.plot(x[outliers], y[outliers], 'ro',
ms=20, mec='red', mfc='red', zorder=1, alpha=0.4)
plt.xlabel('X');
plt.ylabel('Y');
display(Markdown(r"""Without outlier modeling:
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
""".format(*percs_blind.T.ravel())
))
percs_brute = np.percentile(sample[:, :2], [16, 50, 84], axis=0)
display(Markdown(r"""With outlier modeling: 22 parameters
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
* number of outliers: ${6:d}$
""".format(*percs_brute.T.ravel(), n_outliers)
))
Without outlier modeling:
$\alpha$ = 39.7 [38.5, 40.9]
$\beta$ = 0.237 [0.21, 0.264]
With outlier modeling: 22 parameters
$\alpha$ = 31.7 [29.9, 33.5]
$\beta$ = 0.46 [0.419, 0.5]
number of outliers: $3$
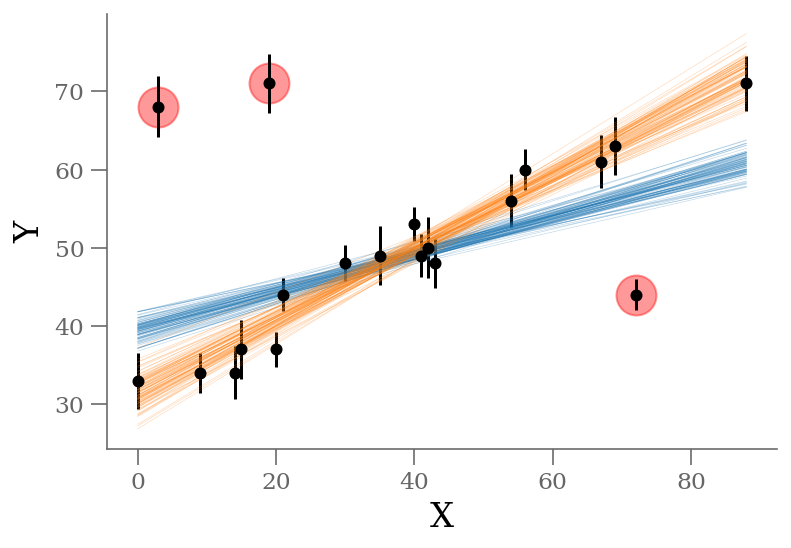
We see a distribution of points near a slope of \(\sim 0.45\), and an intercept of \(\sim 31\). We’ll plot this model over the data below, but first let’s see what other information we can extract from this trace.
One nice feature of analyzing MCMC samples is that the choice of nuisance parameters is completely irrelevant during the sampling.
The result shown in orange matches our intuition. In addition, the points automatically identified as outliers are the ones we would identify by eye to be suspicious. The blue shaded region indicates the previous “blind” result.
The smarter version#
This previous model of outliers takes a simple linear model of \(2\) parameters and transforms it into a \((N+2)\) parameters, \(N\) being the number of datapoints. This leads to \(22\) parameters in our case. What happens if you have \(200\) data points?
The problem with the previous model was that it adds one parameter for each data point, which not only makes the fitting expensive, but also makes the problem untracktable very quickly.
Based on the same formulation, we can consider one global \(q\) instead individuals, which will characterize on the ensemble the probability of having an outlier. In other words, the fraction of outliers relative to the dataset.
with
We simply have \(1\) nuisance parameters: \(q\) which ranges from 0 to 1.
Similarly to the previous model, \(q=0\) indicates an outlier, in which case a Gaussian of width \(\sigma_B\) is used in the computation of the likelihood. This \(\sigma_B\) can also be a nuisance parameter.
From this model, we can estimate the odds of being an outlier with $\( Odds_{outlier}(x_i, y_i, \sigma_i) = \frac{(1-q) P(x_i, y_i | \sigma_{out})}{q\,P(x_i, y_i | \sigma_i, \alpha, \beta) + (1-q) P(x_i, y_i | \sigma_{out})}\)$
Our likelihood can be expressed directly with tensor operations (otherwise we’d need to use theano’s @as_op
decorator). This means we get the likelihood gradient for free thanks to the Theano backend, which makes the MCMC sampler more efficient (HMC methods)
We now make the probabilistic equations that link the various variables together:
We would like to fit a linear model to this above dataset:
One must provide some prior to properly implement the model. Let’s suppose weak informative normal priors (they correspond to a Ridge regression context)
We also have the outlier component \(\hat{y}_{out}\) $\( \hat{y}_{out} \rightsquigarrow \mathcal{N}(\hat{y}_{out}, \sigma_{out})\)\( where \)\sigma_{out}\( is unknown but positively constrained (half-Normal prior) \)\( \sigma_{out} \rightsquigarrow \mathcal{N^+}(0, 100)\)$
\(g\) sets the ratio of inliers to outliers. Given our setup, it corresponds to the fraction of outliers in our model. One can set a wealky informative prior on \(g\) as $\( g \rightsquigarrow \mathcal{U}(0, 0.5) \)$ (hopefully we do not have more than half of the data being made of outliers)
Our final model is a mixture of the two components:
we code the probabilistic model as follow (using DensityDist to set the final mixture likelihood)
def jax_model_outliers(x=None, y=None, sigma_y=None):
## Define weakly informative Normal priors
beta = numpyro.sample("beta", dist.Uniform(0.0, 100))
alpha = numpyro.sample("alpha", dist.Uniform(0.0, 100))
## Define Bernoulli inlier / outlier flags according to
## a hyperprior fraction of outliers, itself constrained
## to [0,.5] for symmetry
frac_outliers = numpyro.sample('frac_outliers',
dist.Uniform(low=0., high=.5))
## variance of outliers
sigma_y_out = numpyro.sample("sigma_y_out", dist.HalfNormal(100))
# linear model
ypred = beta + alpha * x
# inliers
ypred_in = dist.Normal(ypred, sigma_y)
# ouliers
ypred_out = dist.Normal(ypred, sigma_y_out)
with numpyro.plate("i=1..n", len(y), dim=-1):
## define the linear model
p_outlier = numpyro.sample('p_outlier',
dist.Bernoulli(frac_outliers),
infer={'enumerate': 'parallel'})
## need a float for the mixture
probs = jnp.stack([1 - p_outlier,
p_outlier], axis=-1).astype(float)
locs_ = jnp.stack([ypred, ypred], -1)
scales_ = jnp.stack([sigma_y_in * jnp.ones(len(x)),
sigma_y_out * jnp.ones(len(x))], -1)
comp_ = dist.Normal(locs_, scales_)
mixture = dist.MixtureSameFamily(dist.Categorical(probs), comp_)
# calculate the outlier probability for book-keeping
lnl_in = jnp.log(1 - frac_outliers) + ypred_in.log_prob(y)
lnl_out = jnp.log(frac_outliers) + ypred_out.log_prob(y)
ltot = jnp.logaddexp(lnl_in, lnl_out)
q = numpyro.deterministic('q', jnp.exp(lnl_out) / jnp.exp(ltot))
# likelihood
numpyro.sample("y", mixture, obs=y)
numpyro.render_model(jax_model_outliers, model_args=(x, y, sy,),
render_distributions=True)
# Start from this source of randomness. We will split keys for subsequent operations.
rng_key = random.PRNGKey(0)
rng_key, rng_key_ = random.split(rng_key)
# Run NUTS.
kernel = NUTS(jax_model_outliers)
num_samples = 5000
mcmc = MCMC(kernel, num_warmup=3000, num_samples=num_samples)
mcmc.run(rng_key_, x=x, y=y, sigma_y=sy)
mcmc.print_summary()
traces_smart = mcmc.get_samples()
sample: 100%|██████████| 8000/8000 [00:06<00:00, 1190.80it/s, 15 steps of size 3.48e-01. acc. prob=0.93]
mean std median 5.0% 95.0% n_eff r_hat
alpha 0.47 0.04 0.47 0.41 0.53 2428.99 1.00
beta 31.31 1.49 31.30 29.08 33.95 2394.69 1.00
frac_outliers 0.21 0.09 0.20 0.06 0.36 2507.26 1.00
sigma_y_out 41.06 22.91 34.93 14.32 69.36 2038.42 1.00
Number of divergences: 0
samples_q = 1 - traces_smart['q'].T
plt.figure(figsize=(15, 5))
colors = np.where(np.median(samples_q, axis=1) < 0.5, 'C1', 'C0')
fast_violin(samples_q.T, color=colors)
ticks = (np.arange(samples_q.shape[0]) + 1)
lbls = [r'x$_{{{0:d}}}$'.format(k) for k in (0.5 * ticks).astype(int)]
plt.xticks(ticks, lbls)
plt.hlines([0.5], 0.5, len(ticks) + 0.5, color='0.5', zorder=-100, ls=':')
plt.xlim(0.5, len(ticks) + 0.5)
plt.ylabel('q_i');
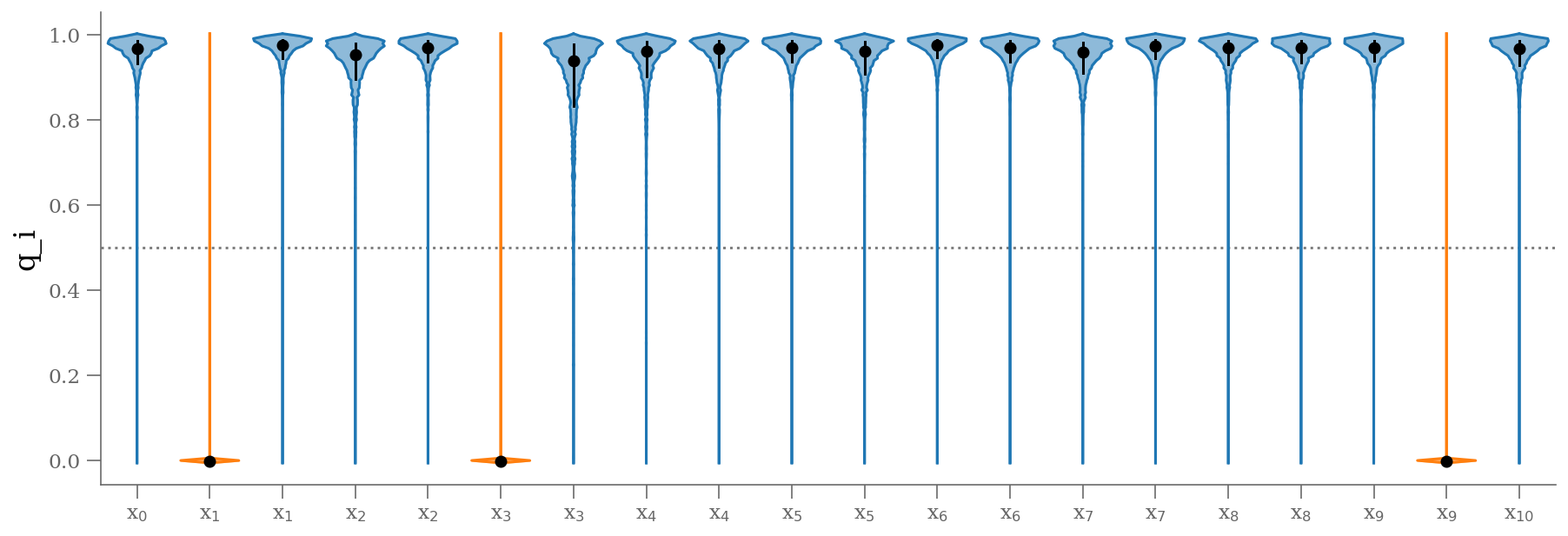
varnames = 'beta', 'alpha', 'frac_outliers', 'sigma_y_out'
sample = np.array([traces_smart[k].ravel() for k in varnames]).T
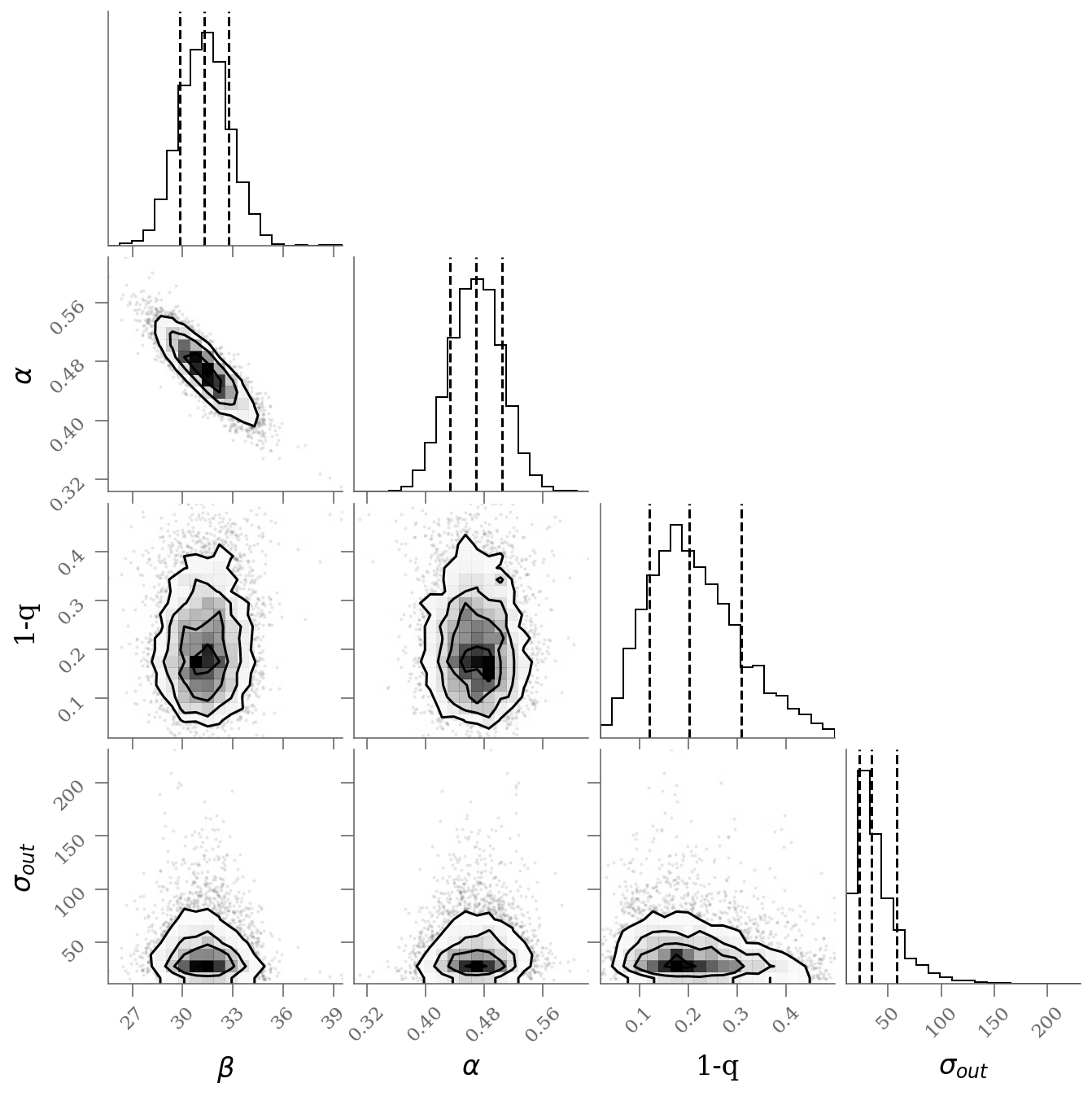
corner(sample[:, :], labels=(r'$\beta$', r'$\alpha$', r'1-q', r'$\sigma_{out}$'), quantiles=(0.16, 0.50, 0.84));
outliers = np.median(traces_smart['q'], axis=0) >= 0.5
n_outliers = int(sum(outliers))
x, y, sy = np.loadtxt('line_outlier.dat', unpack=True)
plt.errorbar(x, y, yerr=sy, marker='o', ls='None', ms=5, color='k');
ypred_smart = sample[:, 0] + sample[:, 1] * x[:, None]
plt.plot(x, ypred_blind[:, :100], color='C0',
rasterized=True, alpha=0.3, lw=0.3, zorder=-3)
plt.plot(x, ypred_brute[:, :1000], color='C1',
rasterized=True, alpha=0.3, lw=0.3, zorder=-2)
plt.plot(x, ypred_smart[:, :1000], color='C2',
rasterized=True, alpha=0.3, lw=0.3, zorder=-1)
plt.plot(x[outliers], y[outliers], 'ro',
ms=20, mec='red', mfc='red', zorder=1, alpha=0.4)
plt.xlabel('X');
plt.ylabel('Y');
display(Markdown(r"""Without outlier modeling (blue):
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
""".format(*percs_blind.T.ravel())
))
display(Markdown(r"""With outlier modeling (orange): 22 parameters
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
* number of outliers: ${6:d}$
""".format(*percs_brute.T.ravel(), n_outliers)
))
percs_smart = np.percentile(sample[:, :-1], [16, 50, 84], axis=0)
display(Markdown(r"""With outlier modeling (green): 3 parameters
* $\alpha$ = {1:0.3g} [{0:0.3g}, {2:0.3g}]
* $\beta$ = {4:0.3g} [{3:0.3g}, {5:0.3g}]
* $q$ = {7:0.3g} [{6:0.3g}, {8:0.3g}]
* number of outliers: ${9:d}$
""".format(*percs_smart.T.ravel(), n_outliers)
))
Without outlier modeling (blue):
$\alpha$ = 39.7 [38.5, 40.9]
$\beta$ = 0.237 [0.21, 0.264]
With outlier modeling (orange): 22 parameters
$\alpha$ = 31.7 [29.9, 33.5]
$\beta$ = 0.46 [0.419, 0.5]
number of outliers: $3$
With outlier modeling (green): 3 parameters
$\alpha$ = 31.3 [29.8, 32.8]
$\beta$ = 0.469 [0.433, 0.504]
$q$ = 0.202 [0.12, 0.308]
number of outliers: $3$
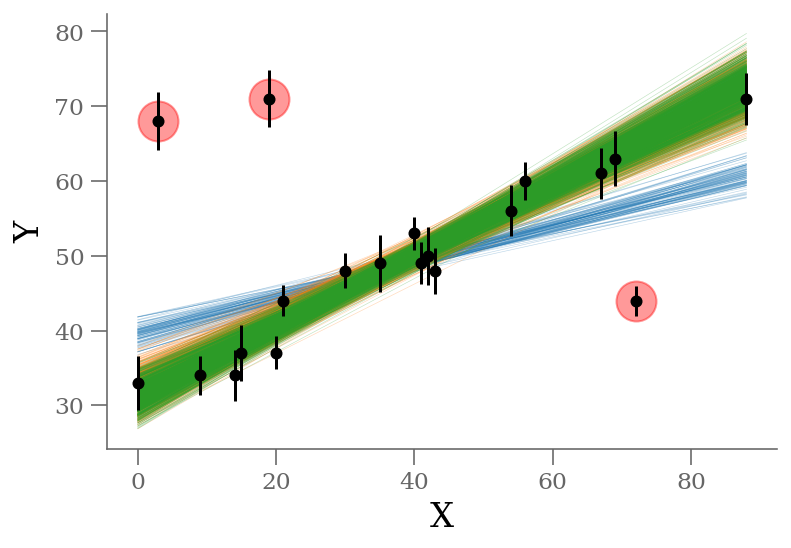
The infered parameters are nearly identical but the convergence is much faster in the last approach.
